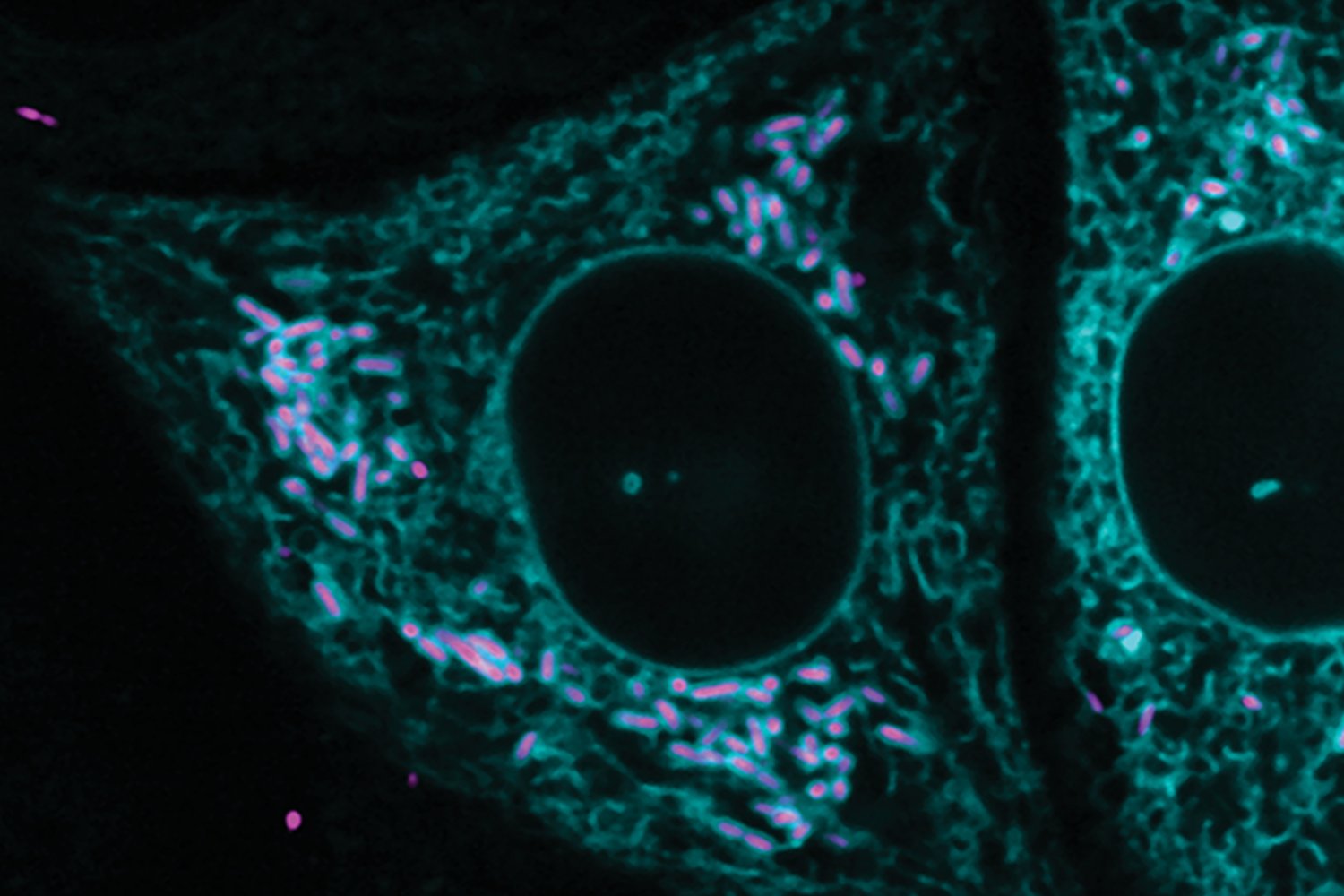
In biology textbooks, the endoplasmic reticulum is often portrayed as a distinct, compact organelle near the nucleus, and is commonly known to be responsible for protein trafficking and secretion. In reality, the ER is vast and dynamic, spread throughout the cell and able to establish contact and communication with and between other organelles. These membrane contacts regulate processes as diverse as fat metabolism, sugar metabolism, and immune responses.
Exploring how pathogens manipulate and hijack essential processes to promote their own life cycles can reveal much about fundamental cellular functions and provide insight into viable treatment options for understudied pathogens.
New research from the Lamason Lab in the Department of Biology at MIT recently published in the Journal of Cell Biology has shown that Rickettsia parkeri, a bacterial pathogen that lives freely in the cytosol, can interact in an extensive and stable way with the rough endoplasmic reticulum, forming previously unseen contacts with the organelle.
It’s the first known example of a direct interkingdom contact site between an intracellular bacterial pathogen and a eukaryotic membrane.
The Lamason Lab studies R. parkeri as a model for infection of the more virulent Rickettsia rickettsii. R. rickettsii, carried and transmitted by ticks, causes Rocky Mountain Spotted Fever. Left untreated, the infection can cause symptoms as severe as organ failure and death.
Rickettsia is difficult to study because it is an obligate pathogen, meaning it can only live and reproduce inside living cells, much like a virus. Researchers must get creative to parse out fundamental questions and molecular players in the R. parkeri life cycle, and much remains unclear about how R. parkeri spreads.
Detour to the junction
First author Yamilex Acevedo-Sánchez, a BSG-MSRP-Bio program alum and a graduate student at the time, stumbled across the ER and R. parkeri interactions while trying to observe Rickettsia reaching a cell junction.
The current model for Rickettsia infection involves R. parkeri spreading cell to cell by traveling to the specialized contact sites between cells and being engulfed by the neighboring cell in order to spread. Listeria monocytogenes, which the Lamason Lab also studies, uses actin tails to forcefully propel itself into a neighboring cell. By contrast, R. parkeri can form an actin tail, but loses it before reaching the cell junction. Somehow, R. parkeri is still able to spread to neighboring cells.
After an MIT seminar about the ER’s lesser-known functions, Acevedo-Sánchez developed a cell line to observe whether Rickettsia might be spreading to neighboring cells by hitching a ride on the ER to reach the cell junction.
Instead, she saw an unexpectedly high percentage of R. parkeri surrounded and enveloped by the ER, at a distance of about 55 nanometers. This distance is significant because membrane contacts for interorganelle communication in eukaryotic cells form connections from 10-80 nanometers wide. The researchers ruled out that what they saw was not an immune response, and the sections of the ER interacting with the R. parkeri were still connected to the wider network of the ER.
“I’m of the mind that if you want to learn new biology, just look at cells,” Acevedo-Sánchez says. “Manipulating the organelle that establishes contact with other organelles could be a great way for a pathogen to gain control during infection.”
The stable connections were unexpected because the ER is constantly breaking and reforming connections, lasting seconds or minutes. It was surprising to see the ER stably associating around the bacteria. As a cytosolic pathogen that exists freely in the cytosol of the cells it infects, it was also unexpected to see R. parkeri surrounded by a membrane at all.
Small margins
Acevedo-Sánchez collaborated with the Center for Nanoscale Systems at Harvard University to view her initial observations at higher resolution using focused ion beam scanning electron microscopy. FIB-SEM involves taking a sample of cells and blasting them with a focused ion beam in order to shave off a section of the block of cells. With each layer, a high-resolution image is taken. The result of this process is a stack of images.
From there, Acevedo-Sánchez marked what different areas of the images were — such as the mitochondria, Rickettsia, or the ER — and a program called ORS Dragonfly, a machine learning program, sorted through the thousand or so images to identify those categories. That information was then used to create 3D models of the samples.
Acevedo-Sánchez noted that less than 5 percent of R. parkeri formed connections with the ER — but small quantities of certain characteristics are known to be critical for R. parkeri infection. R. parkeri can exist in two states: motile, with an actin tail, and nonmotile, without it. In mutants unable to form actin tails, R. parkeri are unable to progress to adjacent cells — but in nonmutants, the percentage of R. parkeri that have tails starts at about 2 percent in early infection and never exceeds 15 percent at the height of it.
The ER only interacts with nonmotile R. parkeri, and those interactions increased 25-fold in mutants that couldn’t form tails.
Creating connections
Co-authors Acevedo-Sánchez, Patrick Woida, and Caroline Anderson also investigated possible ways the connections with the ER are mediated. VAP proteins, which mediate ER interactions with other organelles, are known to be co-opted by other pathogens during infection.
During infection by R. parkeri, VAP proteins were recruited to the bacteria; when VAP proteins were knocked out, the frequency of interactions between R. parkeri and the ER decreased, indicating R. parkeri may be taking advantage of these cellular mechanisms for its own purposes during infection.
Although Acevedo-Sánchez now works as a senior scientist at AbbVie, the Lamason Lab is continuing the work of exploring the molecular players that may be involved, how these interactions are mediated, and whether the contacts affect the host or bacteria’s life cycle.
Senior author and associate professor of biology Rebecca Lamason noted that these potential interactions are particularly interesting because bacteria and mitochondria are thought to have evolved from a common ancestor. The Lamason Lab has been exploring whether R. parkeri could form the same membrane contacts that mitochondria do, although they haven’t proven that yet. So far, R. parkeri is the only cytosolic pathogen that has been observed behaving this way.
“It’s not just bacteria accidentally bumping into the ER. These interactions are extremely stable. The ER is clearly extensively wrapping around the bacterium, and is still connected to the ER network,” Lamason says. “It seems like it has a purpose — what that purpose is remains a mystery.”
